4Department of Computer Science, City University of Hong Kong, Hong Kong, ChinaĪccurate T cell receptor repertoire profiling has provided novel biological and clinical insights in widespread immunological settings however, there is a lack of reference materials in the community that can be used to calibrate and optimize the various experimental systems in different laboratories.3Hematology and Oncology Department, Shenzhen Children's Hospital, Shenzhen, China.1BGI Education Center, University of Chinese Academy of Sciences, Shenzhen, China.The package also provides automated access to HLA alleles frequencies in worldwide human reference populations stored in the Allele Frequency Net Database.Jinghua Wu 1,2 †, Xie Wang 2 †, Liya Lin 2 †, Xuemei Li 2 †, Sixi Liu 3, Wei Zhang 2,4, Lihua Luo 1,2, Ziyun Wan 2, Mingyan Fang 2, Yi Zhao 2, Xiaodong Wang 3, Huirong Mai 3, Xiuli Yuan 3, Feiqiu Wen 3, Changgang Li 3 * and Xiao Liu 2,5 * Converter functions that provide mappings between different HLA naming schemes are based on the MHC restriction ontology (MRO). The package immunotation provides tools for consistent annotation of HLA genes in typical immunoinformatics workflows such as for example the prediction of MHC-presented peptides in different human donors. Reproducible and consistent annotation of HLA alleles in large-scale bioinformatics workflows remains challenging, because the available reference databases and software tools often use different HLA naming schemes. More than 28,000 different HLA alleles have been reported, with significant differences in allele frequencies between human populations worldwide. The repertoire of antigens presented in a given genetic background largely depends on the sequence of the encoded MHC molecules, and thus, in humans, on the highly variable HLA (human leukocyte antigen) genes of the hyperpolymorphic HLA locus. MHC (major histocompatibility complex) molecules are cell surface complexes that present antigens to T cells. retrieve_chain_lookup_table: Retrieve MHC chain lookup table. 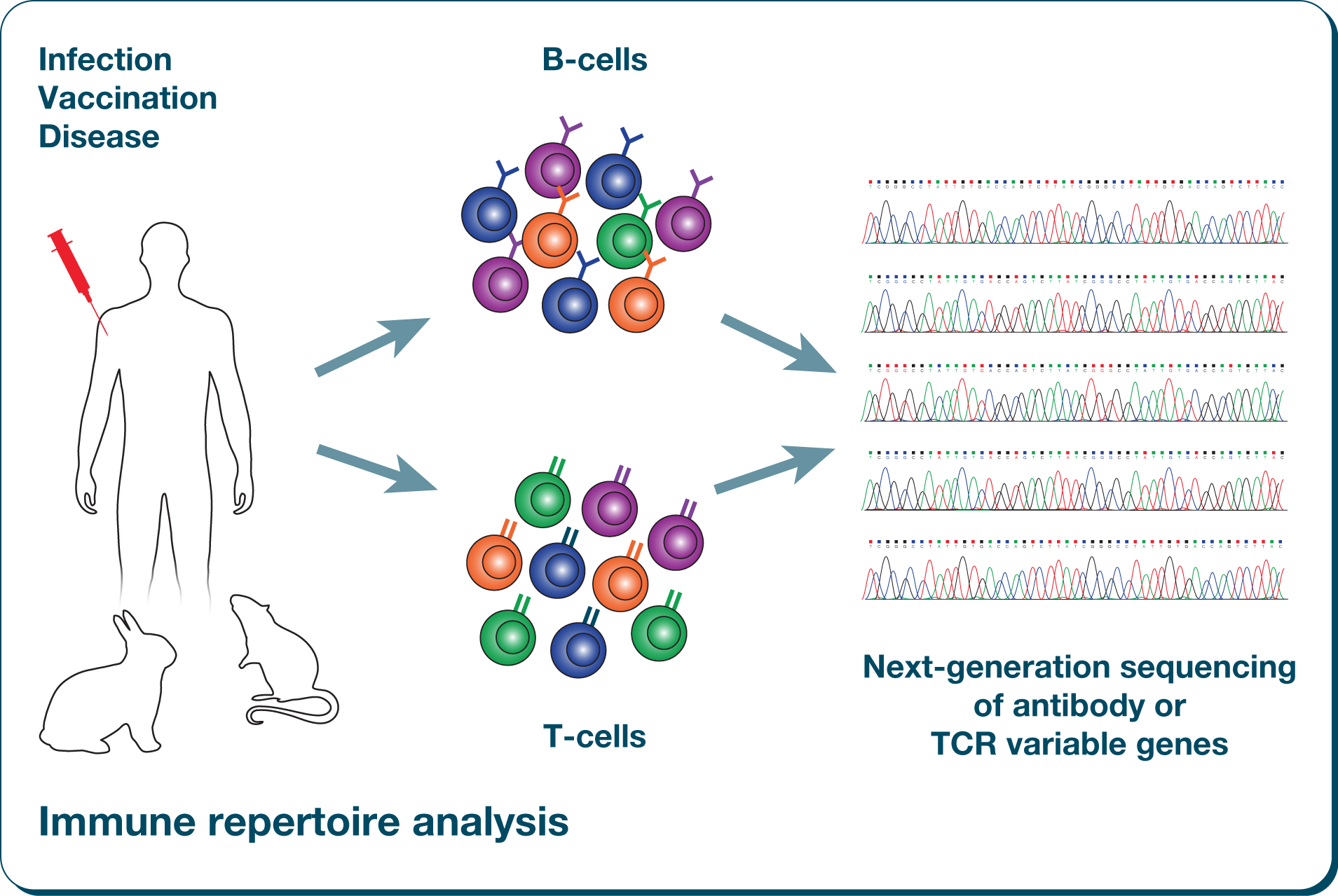
read_population_detail: read_population_detail.read_complete_freq_table: read_complete_freq_table.query_population_detail: Query population metainformation.query_haplotype_frequencies: Query haplotype frequencies.query_allele_frequencies: Query allele frequencies.plot_allele_frequency: Plotting allele frequencies.parse_haplotype_freq_html: parse_haplotype_freq_html.parse_allele_freq_html: parse_allele_freq_html.human_protein_complex_table: human_protein_complex_table.get_valid_organisms: get_valid_organisms.get_valid_geographics: get_valid_geographics.get_mhcpan_input: Get format for NetMHCpan tools.extract_sample_info: extract_sample_info.extract_population_name: extract_population_name. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
